set.seed(5238974)
model_run <- 102
spec_name <- "lookup.yml"
dat.name <- "RawData.csv"
thisScript <- "model_diagnostic_2cmt.qmd"
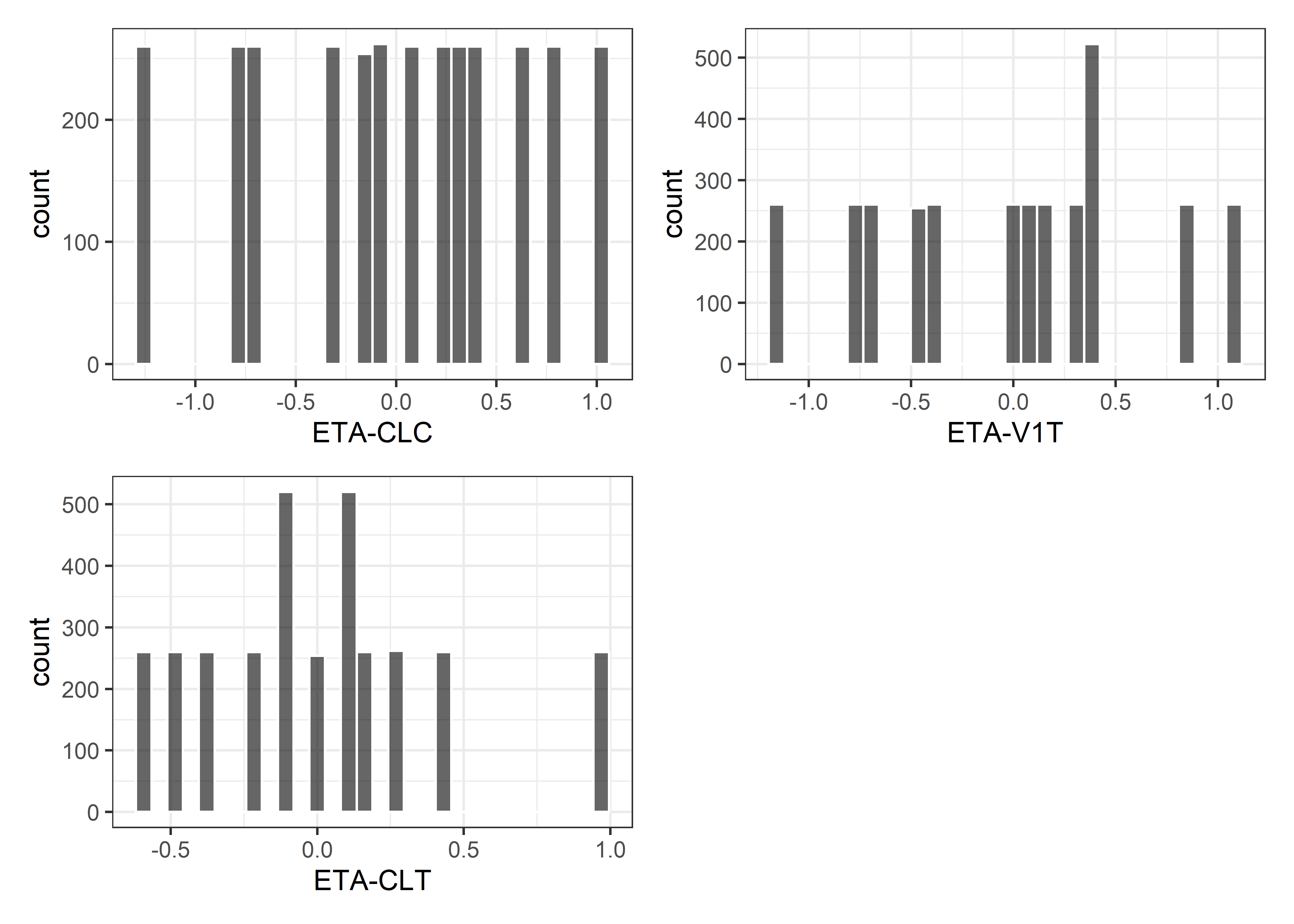
etas <- c("ETA-V1C", "ETA-CLC", "ETA-V2C", "ETA-QC", "ETA-V1T", "ETA-CLT", "ETA-V2T", "ETA-QT")
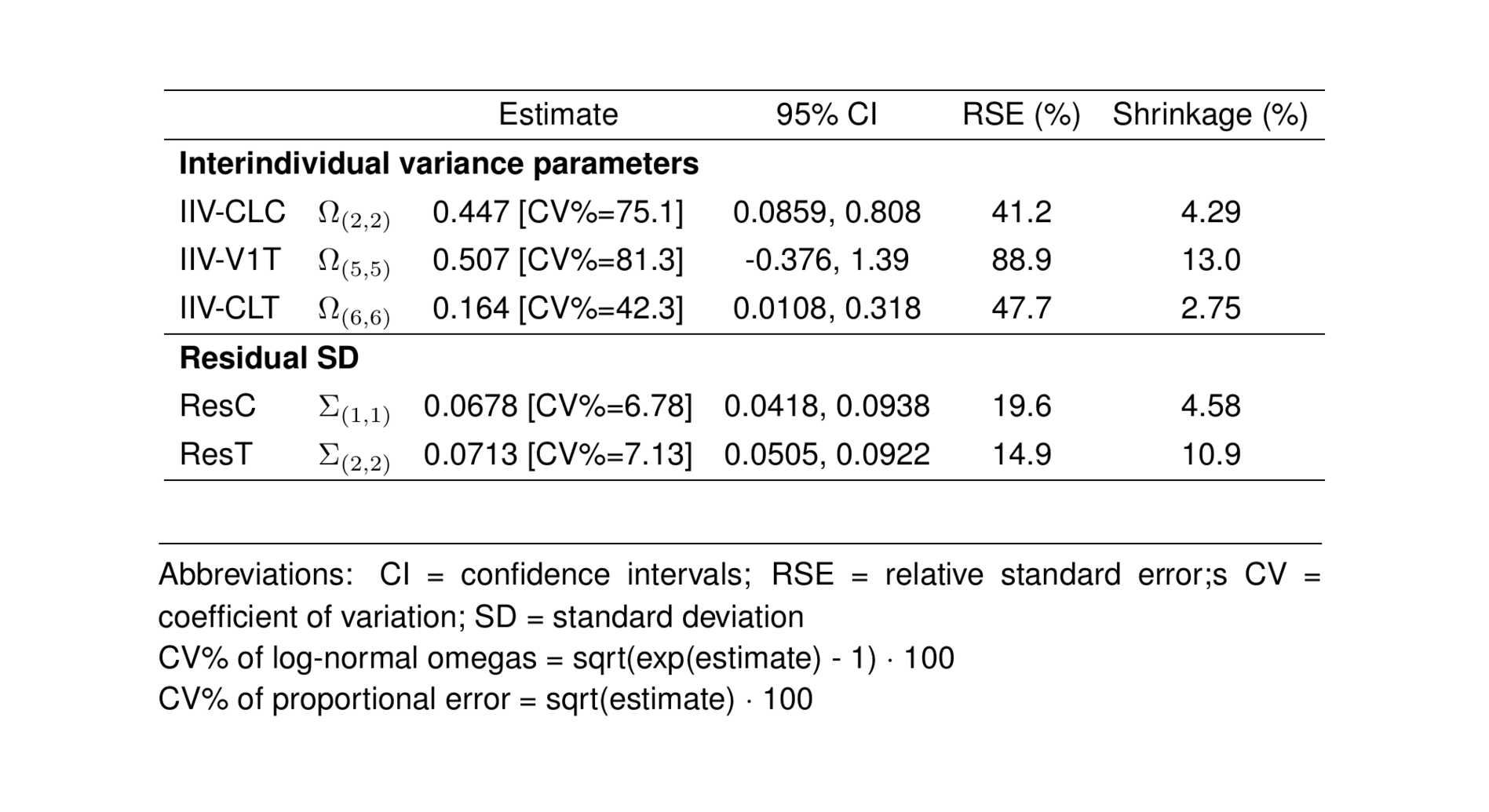
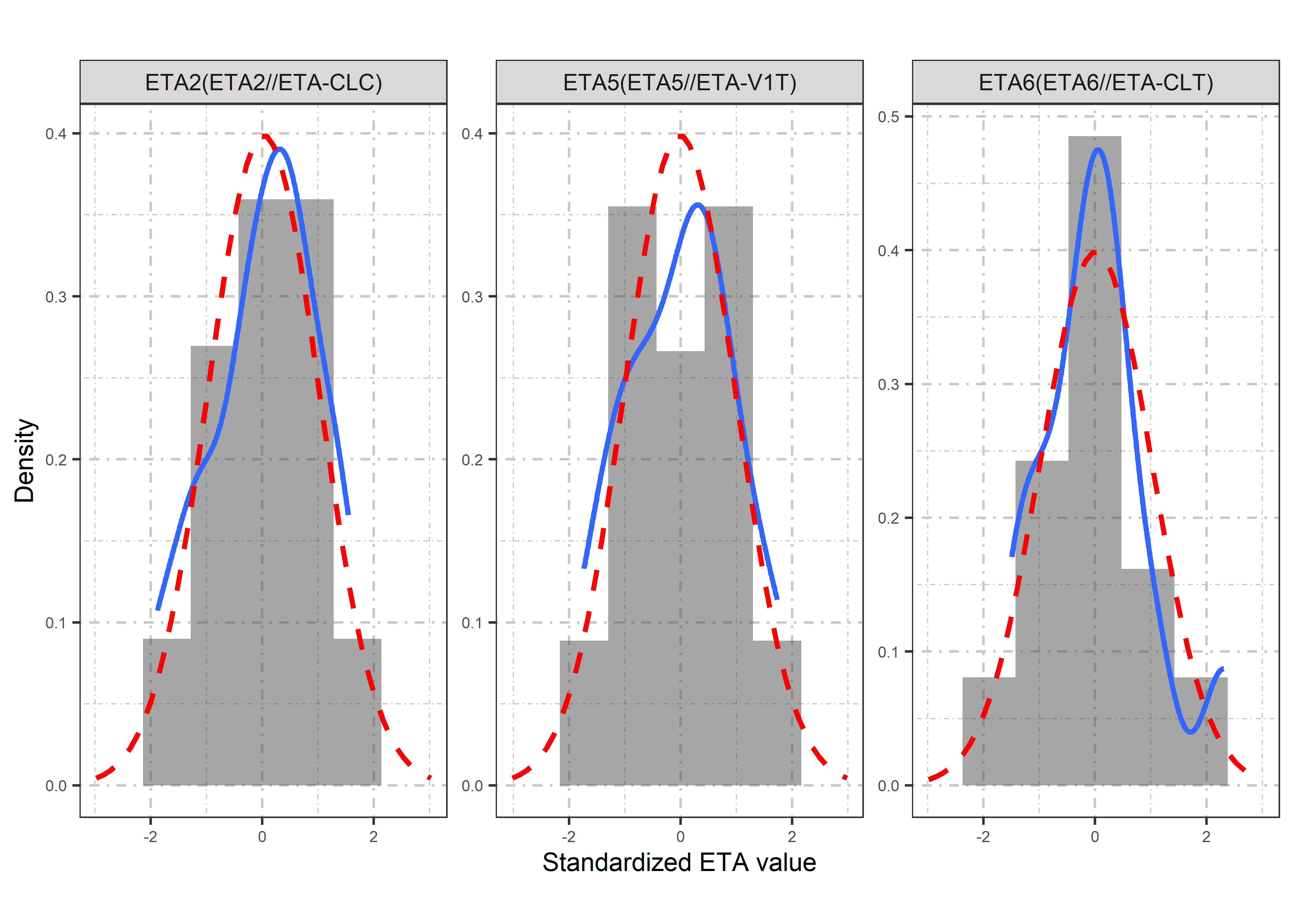
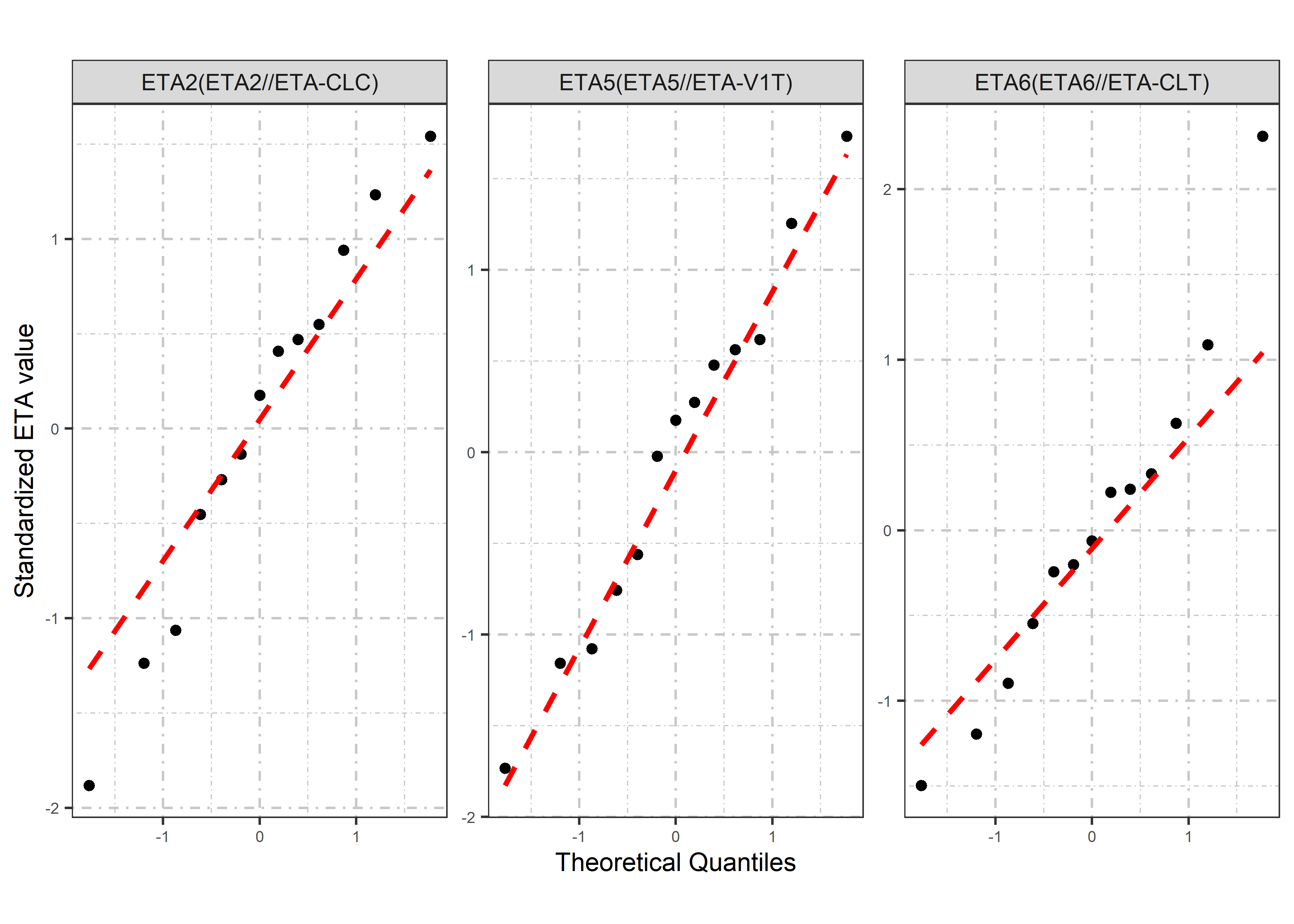
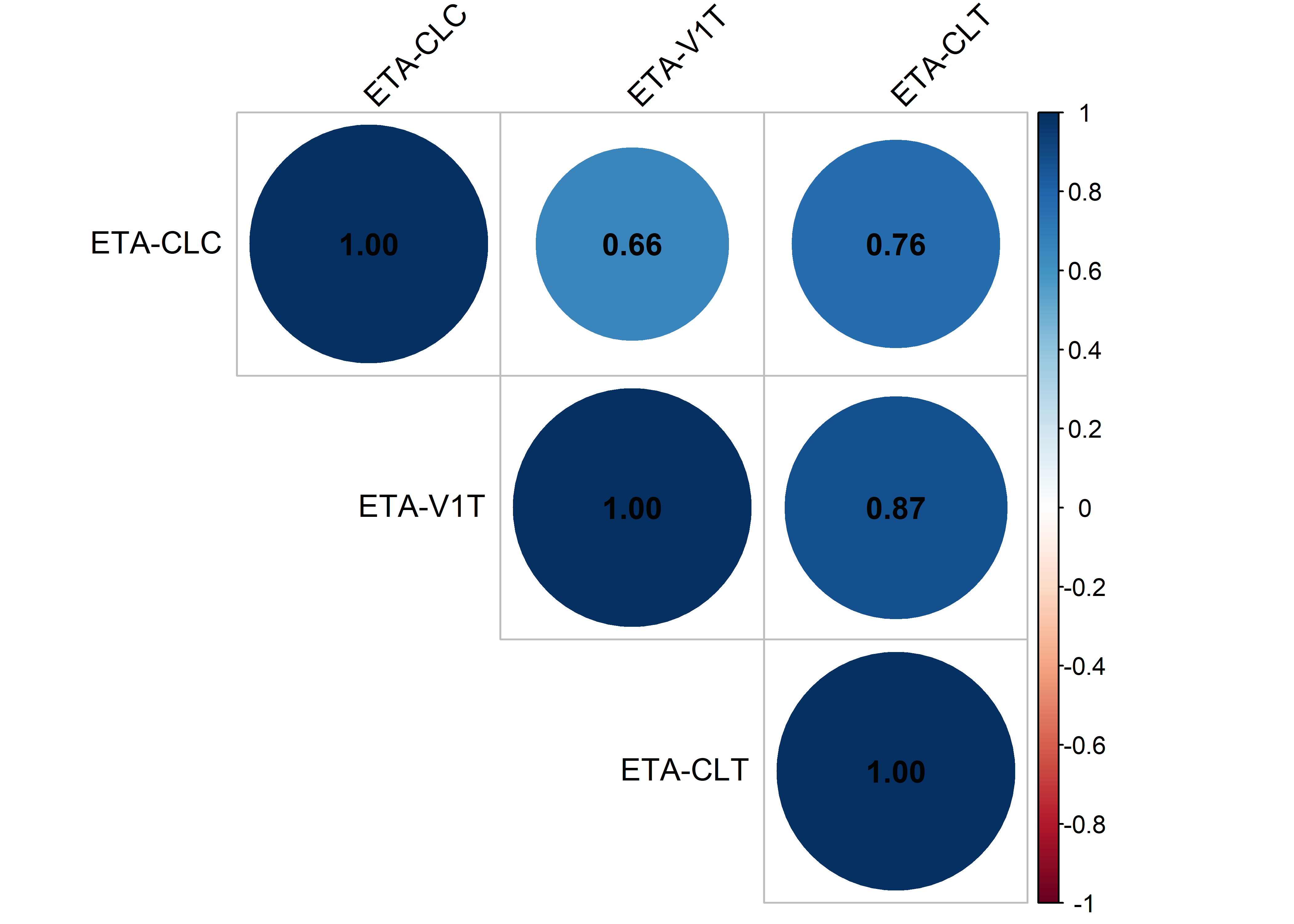
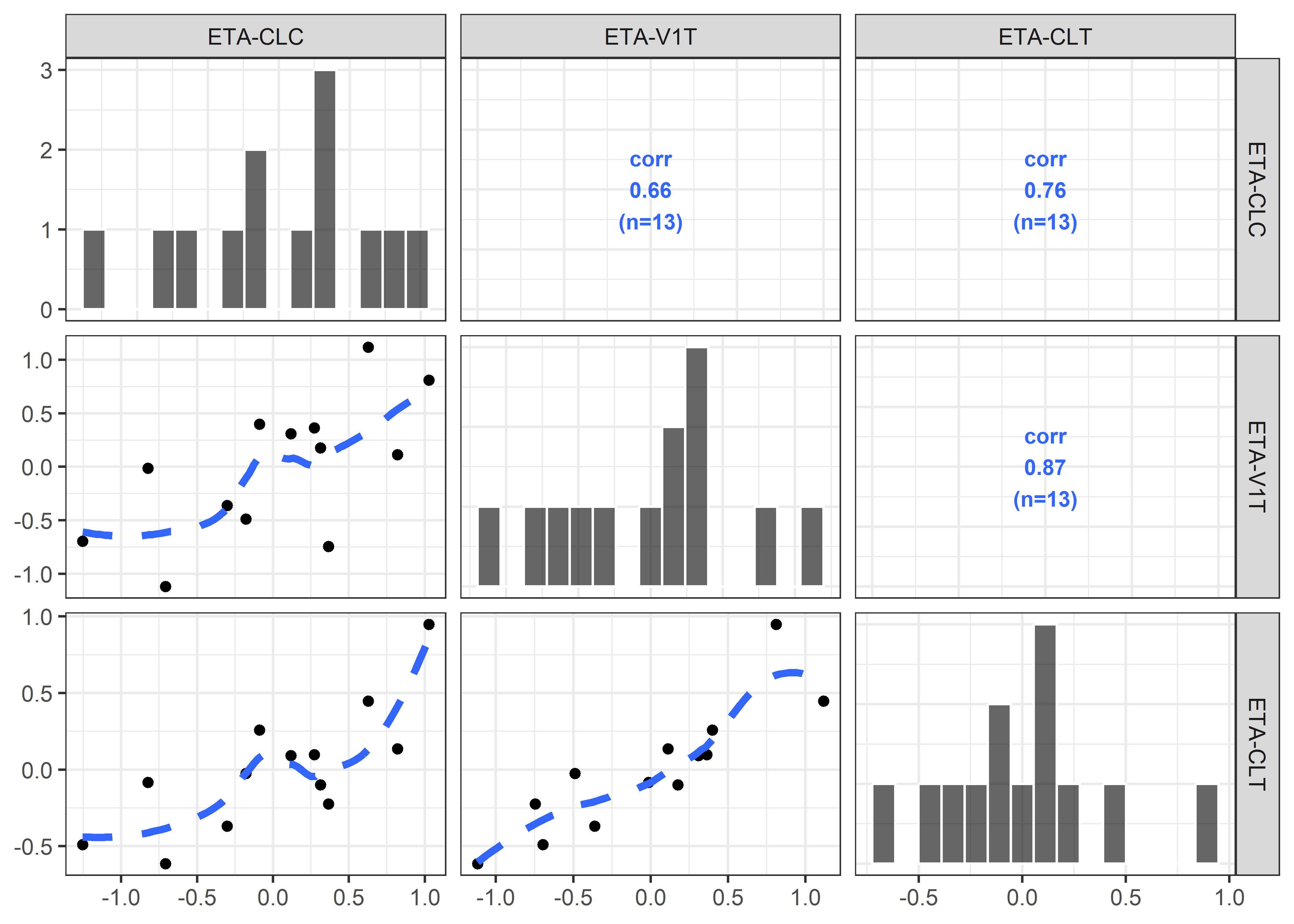
etas_est <- c("ETA2//ETA-CLC", "ETA5//ETA-V1T", "ETA6//ETA-CLT")
#Directories
spec_dir <- here::here("data/derived")
derDir <- here::here("data/derived")
model_dir <- here::here("model/nonmem/basic")
scripts <- here::here("scripts/Model_Development")
tab_dir <- here::here("deliv/tables/Diagnostic")
outDir <- paste0("model/nonmem/basic/", model_run)
options(scipen = 999)
##Continuous Covariates
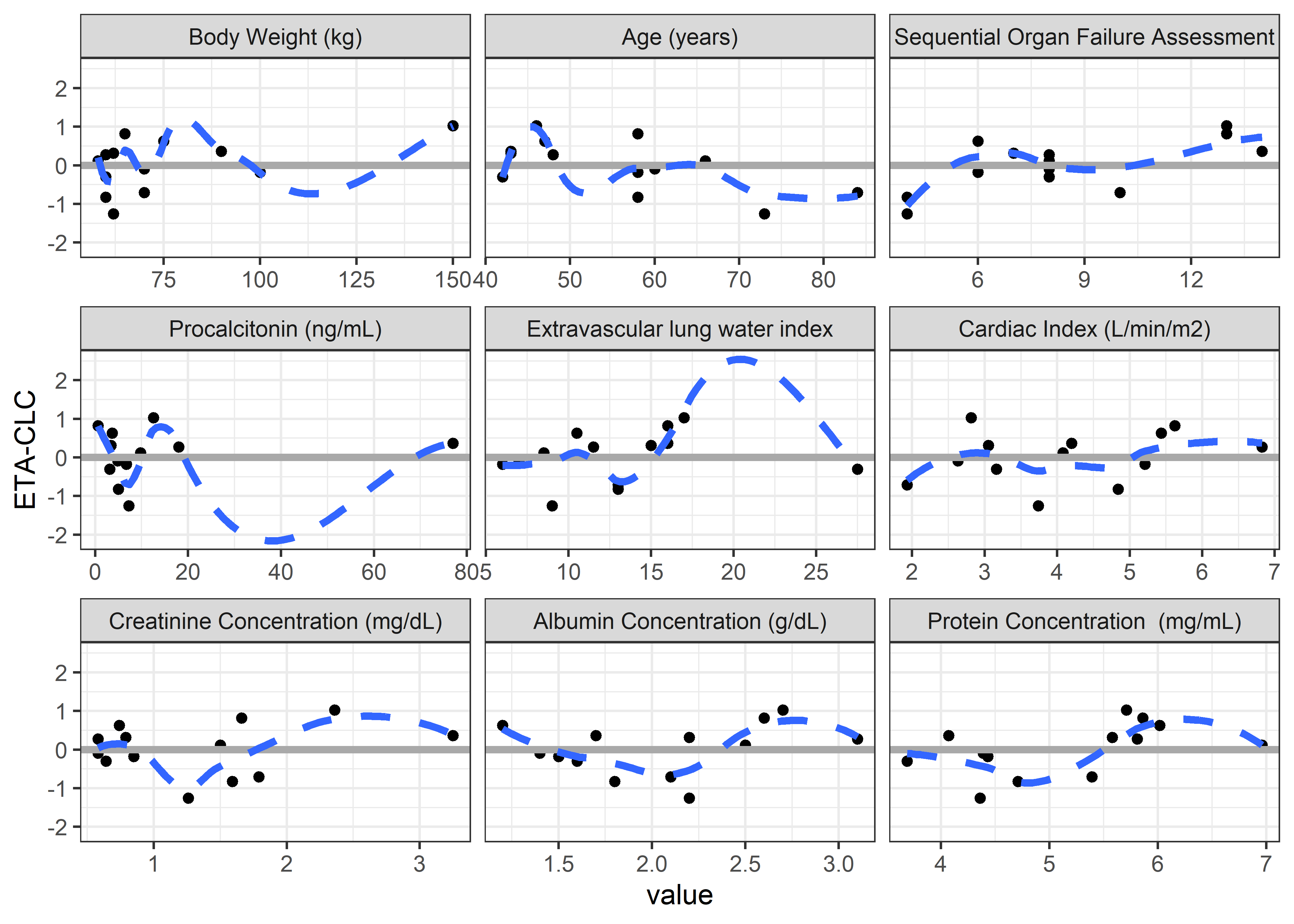
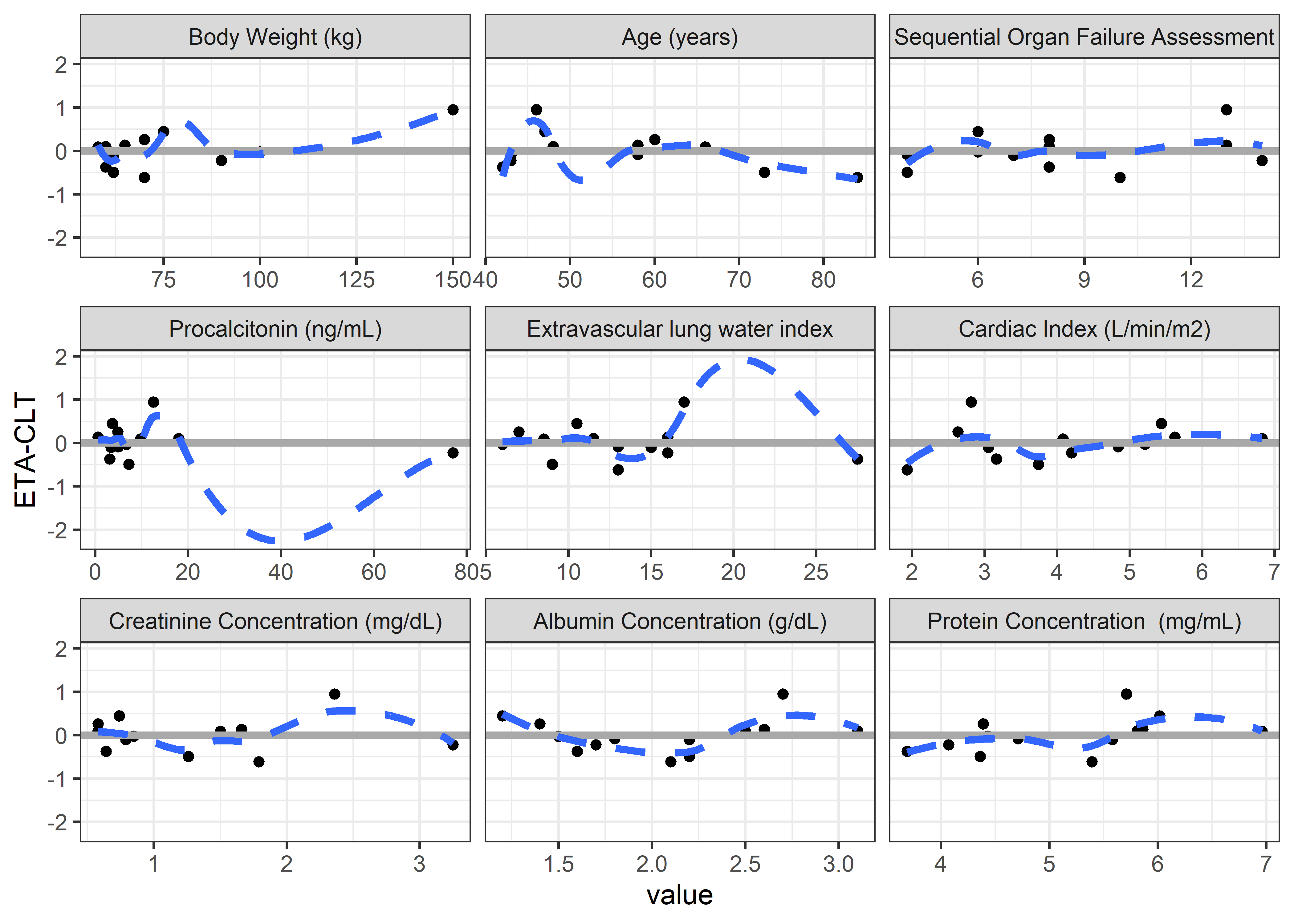
contcov_labels <-c("BW//Body Weight (kg)", "AGE//Age (years)",
"SOFA//Sequential Organ Failure Assessment", "PCT//Procalcitonin (ng/mL)",
"ELWI//Extravascular lung water index", "CI//Cardiac Index (L/min/m2)",
"CRE//Creatinine Concentration (mg/dL)", "ALB//Albumin Concentration (g/dL)",
"PROTEIN//Protein Concentration (mg/mL)") #"HEIGHT//Height (cm)",
contcov_vars <- c("BW", "AGE", "SOFA", "PCT", "ELWI", "CI", "CRE", "ALB", "PROTEIN") #"HEIGHT",
##Categorical Covariates
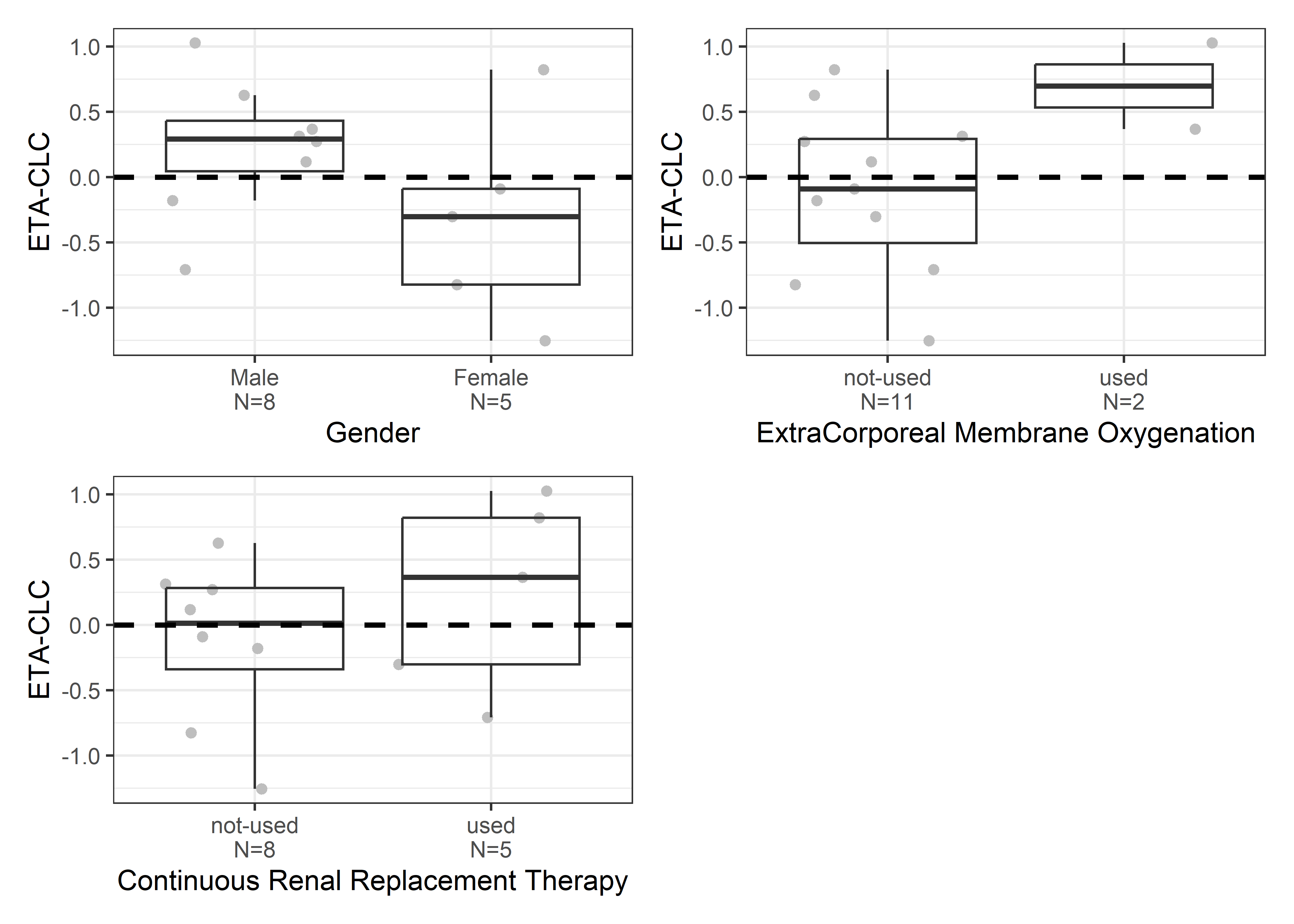
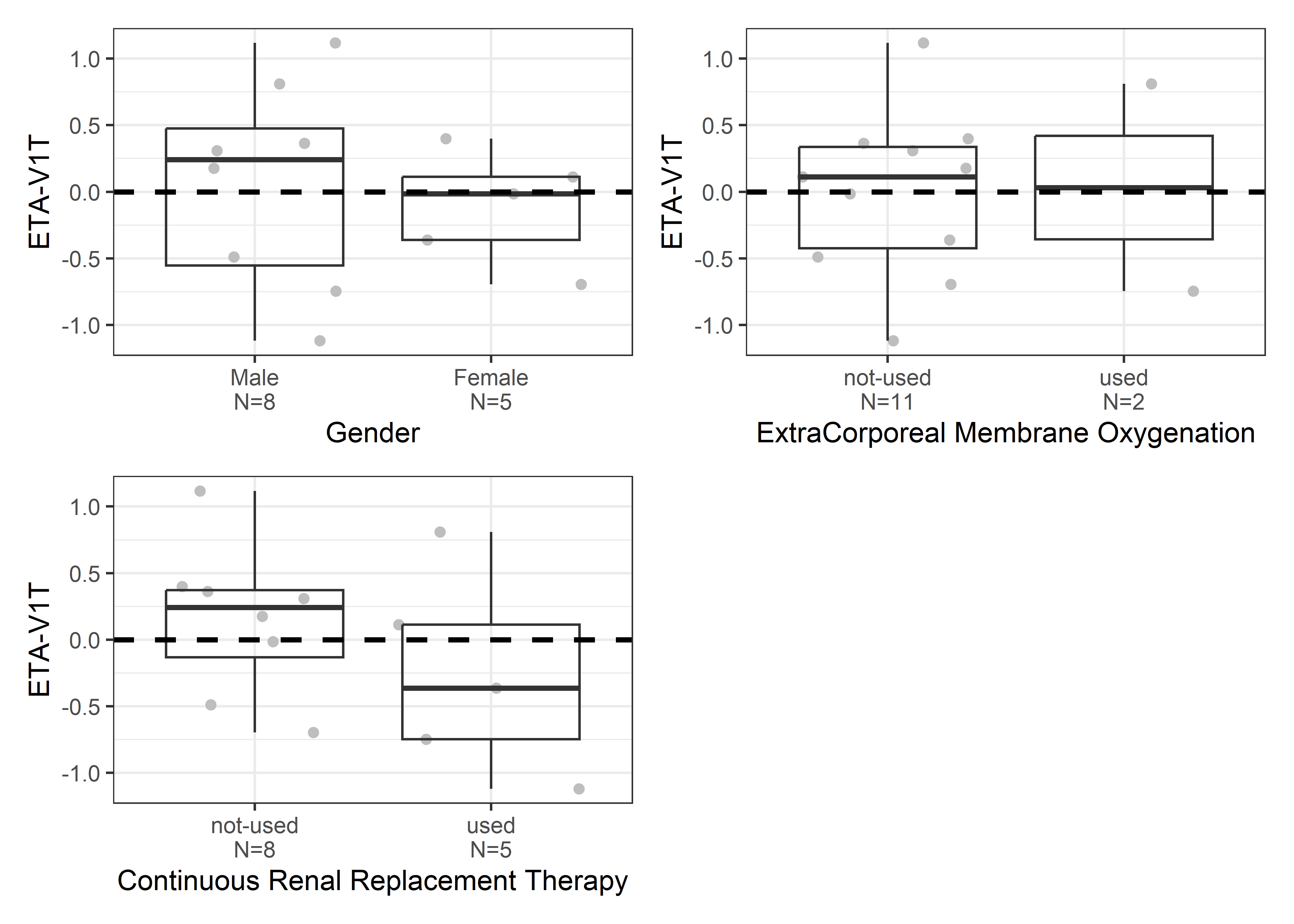
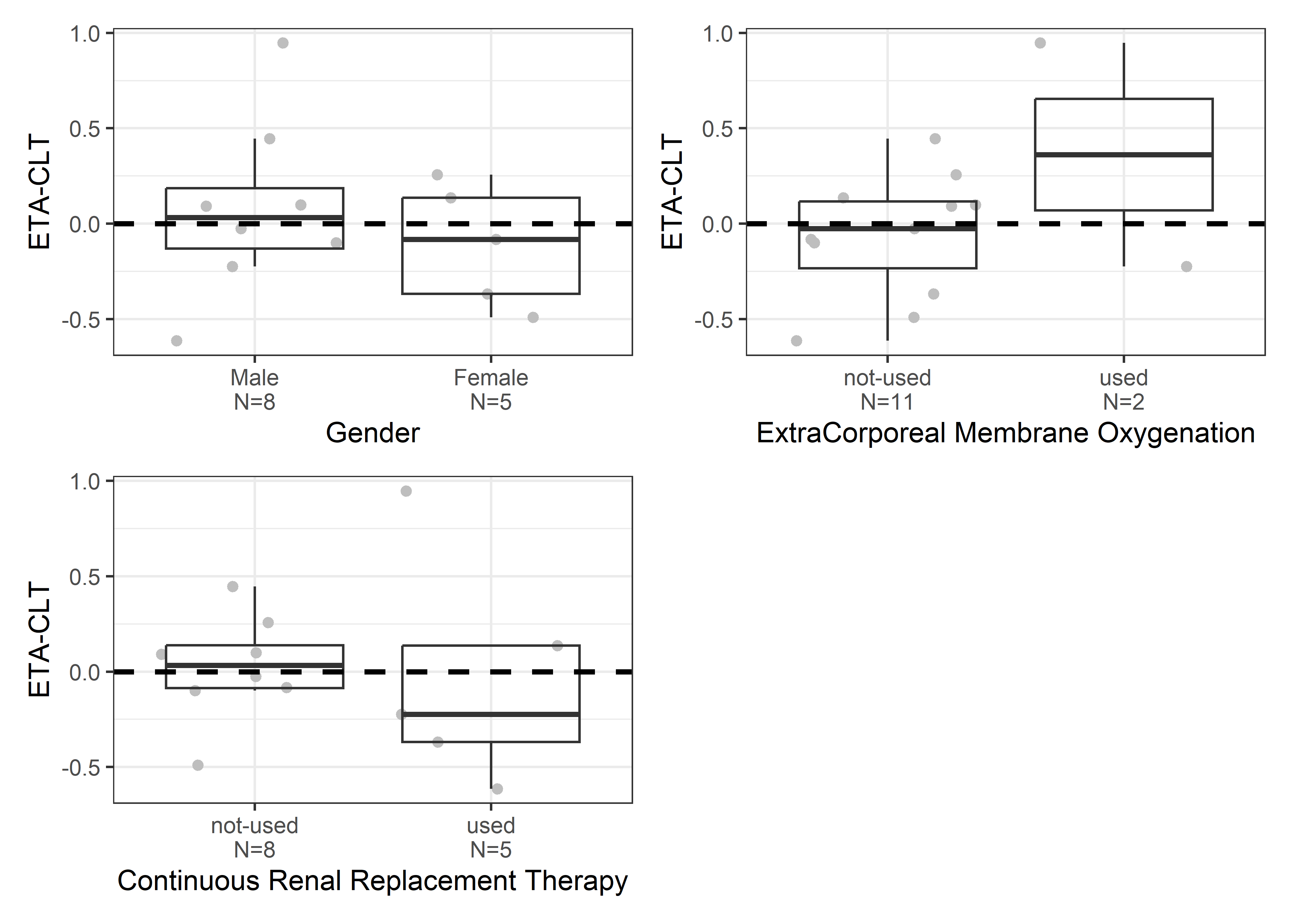
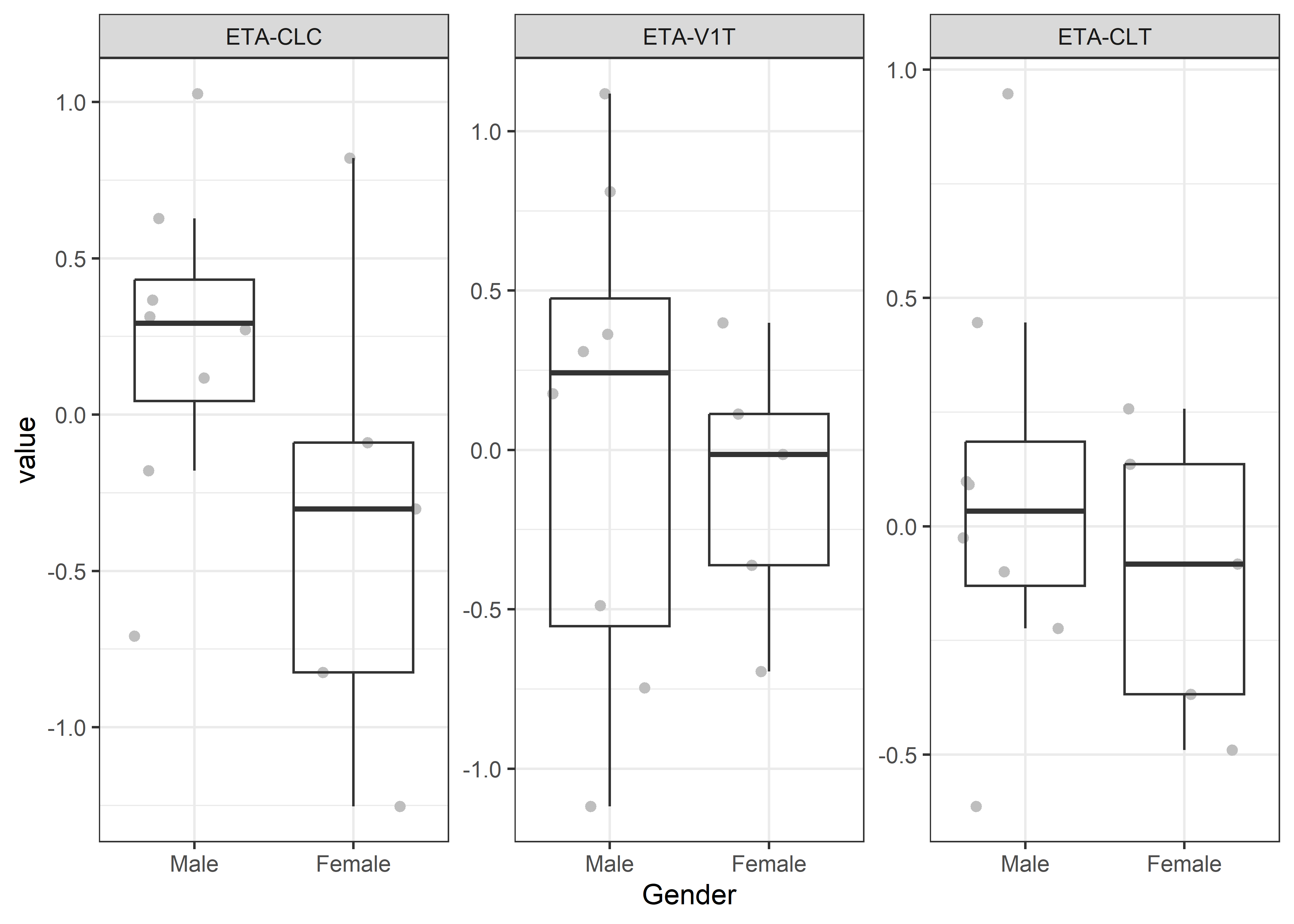
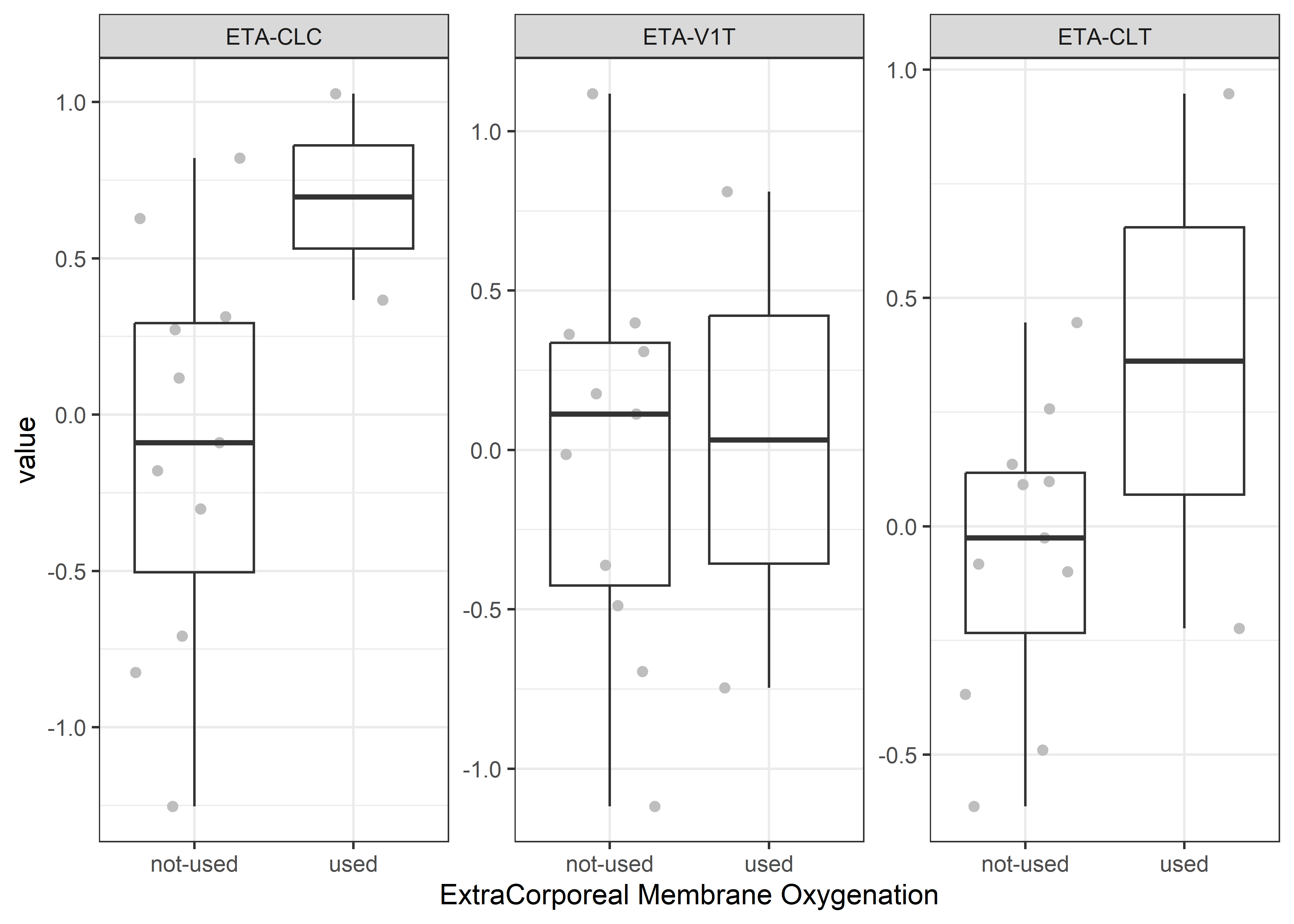
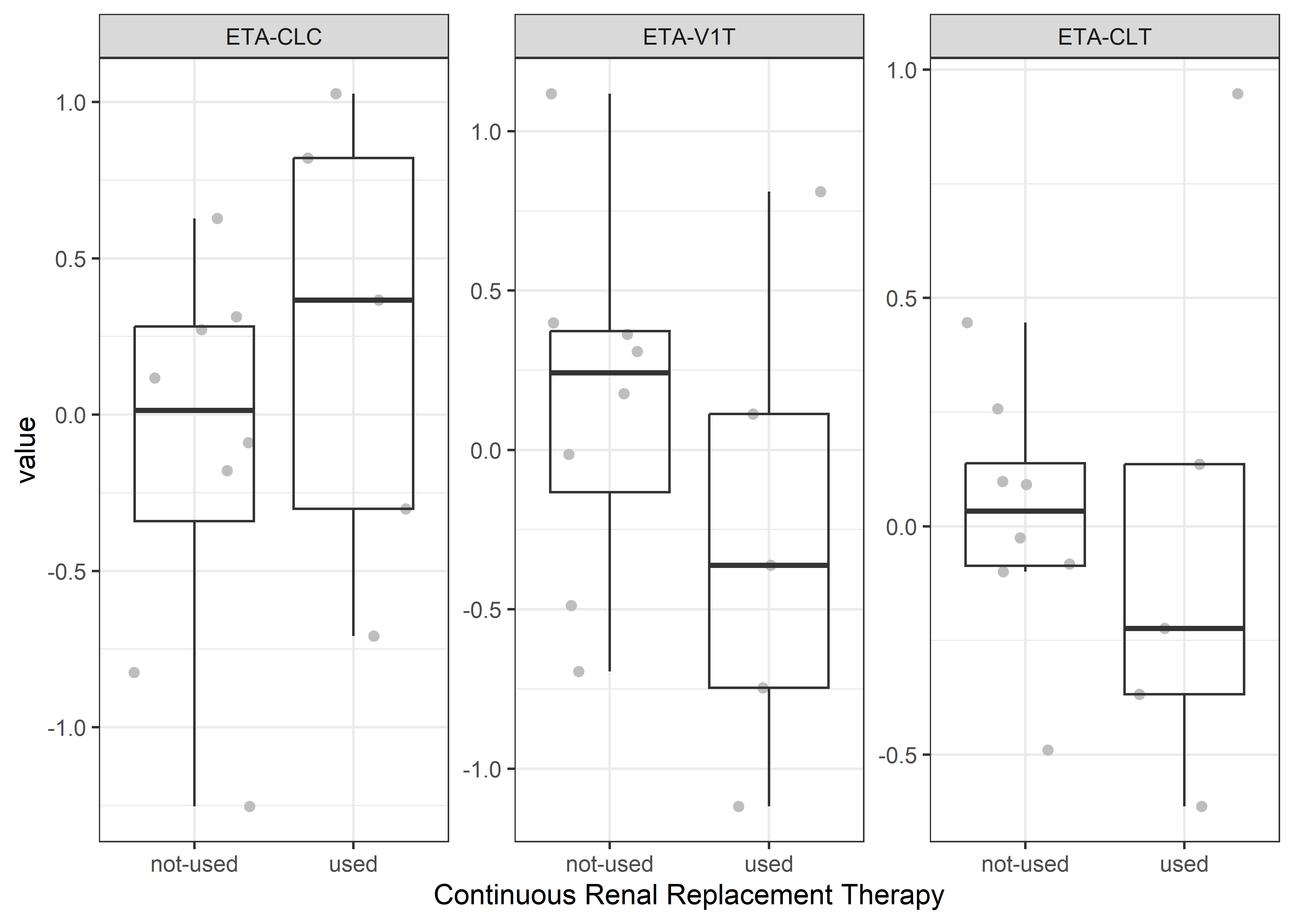
catcov_labels <- c( "SEX_f//Gender", "ECMO_f//ExtraCorporeal Membrane Oxygenation", "CRRT_f//Continuous Renal Replacement Therapy")
catcov_vars <- c("SEX_f", "ECMO_f", "CRRT_f")